DMT-Nexus member
Posts: 58 Joined: 19-Oct-2011 Last visit: 14-Apr-2016 Location: US
|
this whole topic is dumb. why would you transform a vector like yeast to somehow start shitting out DMT when there are plants that already do it. you would still need to purify it some how, and most likely the molecule would be destroyed or kill the organism.
If you wanted to think like a real biologist, you could easily genetically modify the promoter region of whatever the fuck protein(s) make DMT in the PLANT and extract like normally. Or even create a transgenic plant. This of course would cost a shit ton (like grant funded amount of shit ton) and you would need a lab to work in. Shit .... selective breeding would probably be your best route, or engineering a plant utilizing aneuploidy (like strawberries or watermelons)
btw M.S cell/molecular biology here. sorry if i sound like an asshole
|
|
|
|
|
analytical chemist
   
Posts: 7463 Joined: 21-May-2008 Last visit: 03-Mar-2024 Location: the lab
|
yea, that really sounds like the vernacular of an M.S. of Mol Bio  "Nothing is true, everything is permitted." ~ hassan i sabbah
"Experiments are the only means of attaining knowledge at our disposal. The rest is poetry, imagination." -Max Planck
|
|
|
DMT-Nexus member
Posts: 58 Joined: 19-Oct-2011 Last visit: 14-Apr-2016 Location: US
|
lol to that comment
do you know why organisms are transformed to express PROTEIN? DMT is a small molecule, much different than other "drugs" (i guess you could call them that) that could be made in this manor, like insulin, interferon, and other attenuated immunizations.
your going to take an organism that doesn't express the means to produce this chemical AT ALL and somehow make it start by throwing in a few plasmids? please....
find me a scholarly article where something like this has ever even been attempted... then we'll talk.
|
|
|

analytical chemist
   
Posts: 7463 Joined: 21-May-2008 Last visit: 03-Mar-2024 Location: the lab
|
http://www.origene.com/Protein/TP318678.aspxtoo lazy to post the slew of experiments involving expression in e.coli. it's douchebags like you that give grad students a bad name; think you'll come on here and school people  It has been done. Is it practical for amateur practice? probably not. "Nothing is true, everything is permitted." ~ hassan i sabbah
"Experiments are the only means of attaining knowledge at our disposal. The rest is poetry, imagination." -Max Planck
|
|
|

analytical chemist
   
Posts: 7463 Joined: 21-May-2008 Last visit: 03-Mar-2024 Location: the lab
|
Quote:i'm done, troll more bro says the kettle to the pot... you could've just said that this endeavor isn't practical. scientists at a well-known school experimented with cloning d MAW into yeast and expressing the DMAT-synthase pathway, to produce ergolines. why would they do such a dumb experiment? this is analogous to that. "Nothing is true, everything is permitted." ~ hassan i sabbah
"Experiments are the only means of attaining knowledge at our disposal. The rest is poetry, imagination." -Max Planck
|
|
|

Kalt und Heiß, Schwarz und Rot, Kürper und Geist, Liebe und Chaos
 
Posts: 4661 Joined: 02-Jun-2008 Last visit: 30-Apr-2022
|
whitebread420 wrote:this whole topic is dumb. why would you transform a vector like yeast to somehow start shitting out DMT when there are plants that already do it. you would still need to purify it some how, and most likely the molecule would be destroyed or kill the organism. Well, it is not that dumb. there are good reasons to do such a thing; a few of them are: 1. the more dmt sources the better. 2. yeast is something that everyone can grow easily at home. literally everyone, and this is the true key for self-sufficiency (if of course you're after it). There are truly very few people that get their dmt from their garden-grown mimosa or chaliponga or chacruna. Most order these plants. 3. yeast grows rapidly, even phalaris cannot beat it 4. yeast is something very inconspicuous - and cannot be banned by governments! You can order yeast without sweating that the customs may intercept it or without getting ready to make excuses about dyestuff or pot-pourris. whitebread420 wrote:If you wanted to think like a real biologist, you could easily genetically modify the promoter region of whatever the fuck protein(s) make DMT in the PLANT and extract like normally. Or even create a transgenic plant. This of course would cost a shit ton (like grant funded amount of shit ton) and you would need a lab to work in. Shit .... selective breeding would probably be your best route, or engineering a plant utilizing aneuploidy (like strawberries or watermelon Well, genetically modifying the promoter is a grand idea, but you really need the tools to do it in these plants. You got no sequence of the genomic regions you plan to modify neither well-established tools to carry on transfections in such plants. Where would you start from? which loci would you modify? We do not even know the genes that do this kind of job in these plants, we only know the putative enzymatic activities that make dmt in plants. On the contrary, transforming yeast and bacteria is something very very routinely done, it's easy and you do not have to re-invent the wheel. Above all, we have a good idea as to what to do and how too do it (e.g. what genes to add, how to verify them). One could easily write a research proposal for doing such a project - I doubt you'd write a viable proposal with your idea. Creating transgenic plant is a good idea, but again it is not different from making transgenic yeast! Actually it is more difficult and the end-product (i.e. the plant would take much more time to grow, as opposed to the rapid growth of transgenic microbes. Finally, selective breeding is something that takes true dedication and many years - if there's one reason why people go for genetic engineering that's because it's a noce shortcut. Thus far your proposed ideas are not bad, but they do not offer a better alternative to genetically modifying microbes. Since it is a new area, it's better to go from the tried and tested way instead of making things unnecessarily difficult and voluminous. whitebread420 wrote:btw M.S cell/molecular biology here. sorry if i sound like an asshole You do come off a as an asshole, but we forgive you. Just make sure you don't come of as one in the future! Need to calculate between salts and freebases? Click here! Need to calculate freebase or salt percentage at a given pH? Click here!
|
|
|

DMT-Nexus member

Posts: 14191 Joined: 19-Feb-2008 Last visit: 04-Oct-2024 Location: Jungle
|
whitebread420 wrote:this whole topic is dumb. ... shitting out DMT whatever the fuck .... cost a shit ton ... funded amount of shit ton... Shit
btw M.S cell/molecular biology here. sorry if i sound like an asshole If you want to stay in this community, I very very strongly suggest you check the attitude page. This kind of language and attitude really has no place here. Here's an important quote: Attitude page wrote:
- Being aggressive or contentious won’t do anything for you here. You’ll quickly find that people won’t respond to you if they feel your intentions are aggressive or petty.
- Watch your language - Communication is comprised of not only the explicit but also the implicit messages, which are transmited through choice of words and general tone of speech. We do not want curse words and immature slangs in the nexus! Please use language in a dignified manner.
If you want to give constructive criticism, feel free. Calling a topic dumb and using all sorts of curse words and this kind of confrontational attitude will get you nowhere here.
|
|
|
DMT-Nexus member
Posts: 1104 Joined: 17-May-2009 Last visit: 18-Jul-2023
|
What about isolating the nececairy Enzymes from DMT-producing
plants and keeping them alive in some artifical enviroment(petri-dish, Terarria?)
Would that be possible? To take enzymes out of a plant & keep them alive in
an artificial enviroment?
|
|
|

Kalt und Heiß, Schwarz und Rot, Kürper und Geist, Liebe und Chaos
 
Posts: 4661 Joined: 02-Jun-2008 Last visit: 30-Apr-2022
|
SKA wrote:What about isolating the nececairy Enzymes from DMT-producing
plants and keeping them alive in some artifical enviroment(petri-dish, Terarria?)
Would that be possible? To take enzymes out of a plant & keep them alive in
an artificial enviroment?
Ah, if only! enzyme isolation is doable but not that easy and most certainly neither cheap or uncomplicated. Enzymes in dry form and away from heat and light can last for years. But the biggest problem is that for each enzyme to work very particular conditions must be provided, all of these seem far from reach for kitchen chemists. This is why instead of isolating the enzymes, a reasonable strategy is to modify microorganisms to produce them, hoping that these enzymes will perform the desired reactions. Need to calculate between salts and freebases? Click here! Need to calculate freebase or salt percentage at a given pH? Click here!
|
|
|

DMT-Nexus member
Posts: 104 Joined: 28-May-2010 Last visit: 14-May-2023 Location: Earth
|
I'm bringing over some conversation from a thread I started here. It appears that thus far this thread has been limited mainly to discussions of engineering, bacteria/yeast, etc., whereas the topic I'd begun digging into there had mainly to do with those enzymes already identified, kinetic constants and all, in our friend phalaris. Here is one quote from that thread, where I summarize the detailed account of phalaris' enzymatic activity as it is given in this paper. Quote:The decarboxlase in phalaris, with pyridoxal phosphate, yields as much as 1000μMol decarboxylate per mL / per hour. I wouldn’t know how much the enzyme’s observed kinetic constants could matter, I mean, doesn’t this figure tell us everything we need to know already? The methyltransferases in phalaris, with the additional substrate of S-adenosylmethionine (and so with the additional product of S-adenosylhomocysteine) yields as much as 2200μMol methylate per mL / per hour. Again, the enzyme’s observed kinetics are available, but what else than this would we need to know, other than maybe ratios of dimethylations to methylations we could expect to pull at different stages in the reaction? Given these figures, and assuming there is some way of flushing the products from the enzymes (microfilters and a vacuum could work for this, no?) as well as then add additional substrate, I don’t understand why enzymatic synthesis isn’t more commonplace. I mean, all the info on their extraction/separation that I can find would seem to suggest processes not too much more complicated than those for alkaloids.
|
|
|

DMT-Nexus member
Posts: 104 Joined: 28-May-2010 Last visit: 14-May-2023 Location: Earth
|
Okay, so benzyme and Infundibulum both seem to concur that enzyme extraction/isolation is off the table for anyone without access to lab.
So... What about micropropagation? Is this too, much too difficult? I'm thinking here of something along the lines of a jimmy-rigged bioreactor... Where friable callus of phalaris (or whatever of our favorite plants) would be used to initiate a liquid cell suspension culture within a bioreactor: feeding pump would be media rich with L-Tryptophan, effluent would be media rich with the its metabolites?
|
|
|

analytical chemist
   
Posts: 7463 Joined: 21-May-2008 Last visit: 03-Mar-2024 Location: the lab
|
tryptophan decarboxylation seems to be a rate-limiting step in the process of substituted-tryptamine biosynthesis, thus, many organisms produce their own tryptophan, and adding more would be counterproductive. More favorable precursors would be N-Methyltryptamine, or tryptamine. "Nothing is true, everything is permitted." ~ hassan i sabbah
"Experiments are the only means of attaining knowledge at our disposal. The rest is poetry, imagination." -Max Planck
|
|
|

DMT-Nexus member
Posts: 104 Joined: 28-May-2010 Last visit: 14-May-2023 Location: Earth
|
According to those guys, DMT is the rate limiting factor for decarboxlyase and both N-methyltransferases in phalaris. Although, I see you are correct that the tryptophan is endogenous, which I guess would imply, for the experiment being suggested here, we'd want a medium rich with auxins of one type or another instead, right?
|
|
|

analytical chemist
   
Posts: 7463 Joined: 21-May-2008 Last visit: 03-Mar-2024 Location: the lab
|
looking at Scheme 1., it reinforces what I already said. Decarboxylation of TRP is the rate limiting step, and auxin doesn't fit anywhere in that pathway. Like I said, and the paper implies, add tryptamine, or NMT. this would effectively inactivate TDase via feedback inhibition. "Nothing is true, everything is permitted." ~ hassan i sabbah
"Experiments are the only means of attaining knowledge at our disposal. The rest is poetry, imagination." -Max Planck
|
|
|

Barry
Posts: 1740 Joined: 10-Jan-2010 Last visit: 05-Mar-2014 Location: Inside the Higgs Boson
|
Yeast containing dmt mmm. Then make beer add some caapi and you gots yourself ayahuasca in a can  I scare myself with how scientifically clever iam 
|
|
|
member for the trees
  
Posts: 4003 Joined: 28-Jun-2011 Last visit: 27-May-2024
|
..in [ 5,6-dibromo-N,N-DMT - the undersea equation] i suggested genetically tampered with yoghurt to produce DMT (as a bacteria was implicated in making 5bromoDMT in sea-sponges) ..then Tordyveln posted this YouTube video on how to genetically engineer yoghurt at home..! - [ Hacking Youghurt - Tuur van Balen] ..i notice in [ Lactic acid bacteria profiles and tyramine and tryptamine contents of Turkish tulum cheeses] that tyrosine, tryptophan, tyramine & tryptamine are found in cheese making cultures of Lactic Acid Bacteria..as milk naturally contains tryptophan, presumably the bacteria are turning it (&/or tyramine?) into tryptamine.. just patch in another gene or two for DMT..  .
|
|
|

analytical chemist
   
Posts: 7463 Joined: 21-May-2008 Last visit: 03-Mar-2024 Location: the lab
|
welp, start saving your pennies...about 900,000 or so, give or take a few thousand. easy on paper. "Nothing is true, everything is permitted." ~ hassan i sabbah
"Experiments are the only means of attaining knowledge at our disposal. The rest is poetry, imagination." -Max Planck
|
|
|
member for the trees
  
Posts: 4003 Joined: 28-Jun-2011 Last visit: 27-May-2024
|
..well, being a dreamer has it's pleasant moments..
T-Yoghut Consortium anyone..?
|
|
|

DMT-Nexus member
Posts: 104 Joined: 28-May-2010 Last visit: 14-May-2023 Location: Earth
|
benzyme wrote:looking at Scheme 1., it reinforces what I already said. Decarboxylation of TRP is the rate limiting step, and auxin doesn't fit anywhere in that pathway. Like I said, and the paper implies, add tryptamine, or NMT. this would effectively inactivate TDase via feedback inhibition.
I think I just wanted to namedrop "auxin"  The first time I saw the skeletal notation for IAA, I was amazed at what a cousin it was, I mean, it must be that these indoles are what our friends are making tryptophan, etc., from, no? But anyways... Setting aside the actual kinetics for a moment, it would be fairly simple to make a try at something like this wouldn't it? Micropropagation? Liquid cell suspension? What do you think our ideal nutrient medium would look like here? Whitebread420 already made the point above that the whole thread is pretty silly considering that plants already do all of this anyways without any need for mucking about. But the thing is, if a liquid cell suspension could be achieved, and some sort of simple bioreactor imagined, we'd at least be getting more in the ball park of being able to sway towards substitutions most desired by substrates introduced, don't you think? Ultrafiltraters are already in range of a hundred dollars or so... But then I'll be damned if the greatest thing still wouldn't be having each enzyme on its own (and by the GRAM!)... I can't believe it's 1000s and 1000s of dollars for mere micrograms of these things... And have you read about this stuff? Where enzymes are immobilized actively into PVP mesh-arrays, and the substrate can simply be "pumped through" this mesh while desired product comes out the other end? It really is something to dream on for someone with all the aspirations of an organic chemist, but without the stomach for monkeying with "sodium cyanoborohydride." EDIT: Benzyme, you've made it quite clear how impractical enzyme expression is outside the lab... But why is it that in most fancy labs today they don't use enzymes in the way we're talking about here for syntheses? Why isn't every million dollar lab doing enzyme catalyzed reactions along these lines already? I mean, couldn't they replicate these things by kilogram and streamline so many other costs of research/production?
|
|
|

analytical chemist
   
Posts: 7463 Joined: 21-May-2008 Last visit: 03-Mar-2024 Location: the lab
|
whitebread was merely suggesting what infundibulum and I had already pointed out...it's not a cheap nor practical project. Paying for all the equipment and reagents is half of it. The other half is getting the proteins (enzymes) of interest to express in vitro. Of course, the organism should already express proteins for tryptophan metabolism (and many bacterial and yeast cells do); keep in mind, these proteins are substrate specific, so you can't toss in any molecule, like Indole-3-Acetic acid, into a vat of cells expressing INMT, and expect it to get converted to DMT. it doesn't work like that. 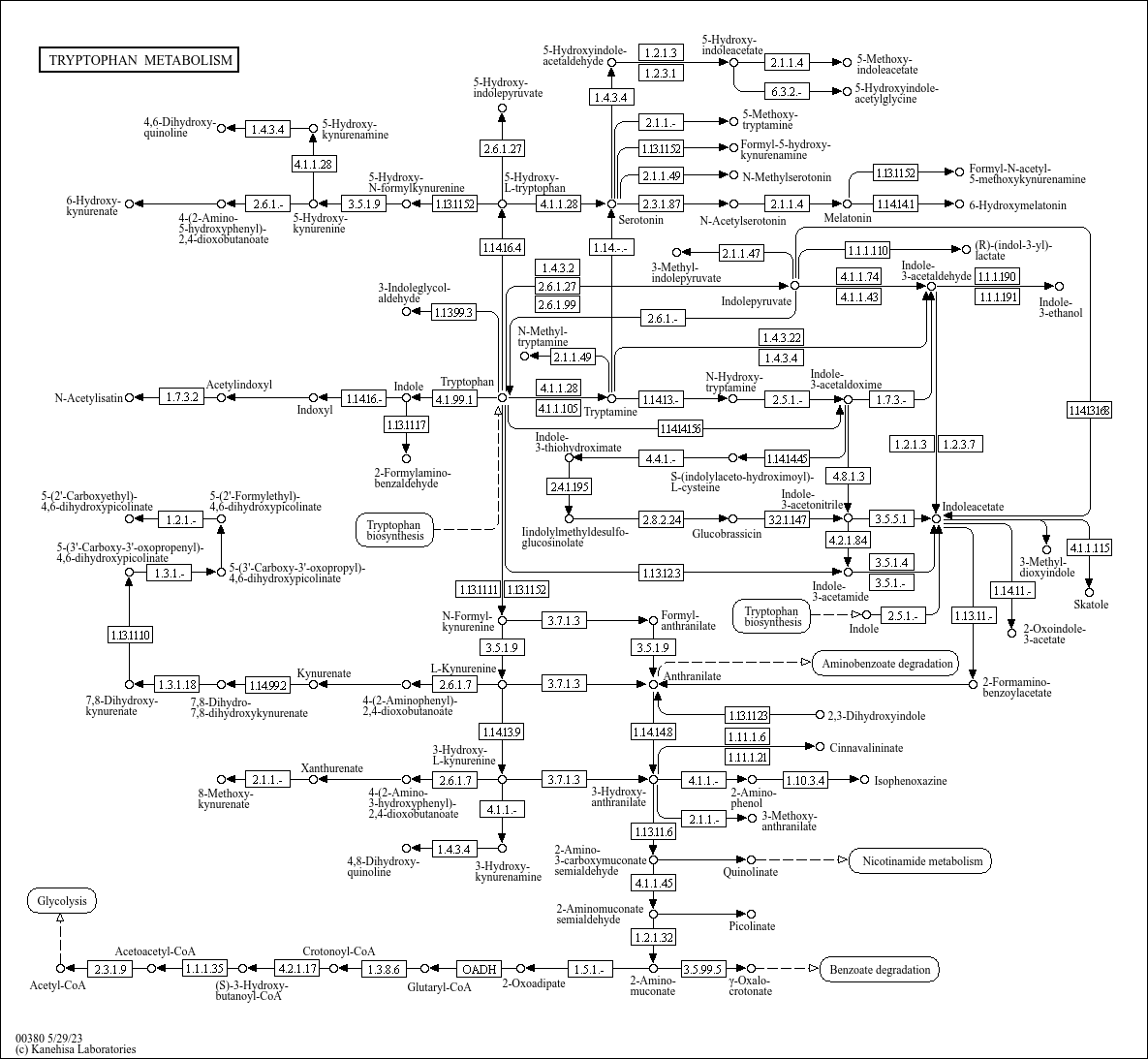 the straightforward route is tryptophan -4.1.1.28-> tryptamine + SAM -2.1.1.49-> NMT + SAM -2.1.1.49-> DMT "Nothing is true, everything is permitted." ~ hassan i sabbah
"Experiments are the only means of attaining knowledge at our disposal. The rest is poetry, imagination." -Max Planck
|